Transfer Learning for TEM Image Segmentation
- Abstract number
- 185
- Presentation Form
- Contributed Talk
- DOI
- 10.22443/rms.mmc2023.185
- Corresponding Email
- [email protected]
- Session
- Artificial Intelligence
- Authors
- Natalia da Silva de Sa (2), Dr. Andrew Stewart (1)
- Affiliations
-
1. University College London
2. University of Limerick
- Keywords
TEM, Transfer Learning, Image Segmentation
- Abstract text
Summary
As a new approach to overcome the shortage of labelled data in the field of Transmission Electron Microscopy (TEM), we investigate the use and applicability of Transfer Learning algorithms for particle segmentation within TEM images. By replacing the encoder of the U-Net architecture with a pre-trained MobileNet, we have found improvements of image segmentation results.
Introduction
Transmission Electron Microscopes (TEM) currently offers such a high data rate that more sophisticated methods of analysis are required, once the data volume is becoming too high to manually shift through images. Manual identification of particles, currently used by the majority of researchers, can be inefficient in terms of time, accuracy and reproducibility of data analysis, because only a small subset of the outputted data is analysed in detail. Machine Learning algorithms for image analysis have demonstrated rapid and significant improvement of the algorithm's ability to accurately segment data, with applications on various research fields, such as autonomous vehicles [2] and object classification using large datasets, such as ImageNet [3], classifying objects with more than 90% accuracy [4].
In contrast, the TEM field faces a shortage of data curated for machine learning applications. We therefore investigate the use of Transfer Learning approaches applied to TEM image data for segmentation. Transfer Learning is a methodology in machine learning which uses a pre-trained network and applies the pretrained network to a second separate dataset using one of the two main approaches: fine-tunning or feature extraction. Feature extraction is most commonly used for datasets with less then 1000 images [5] and it is distinguished by the fact that the pre-trained network is frozen during training and a second network, added to the end of the first, will be trained on a small sample of labelled data, to segment the new data type.
Methods
Applying transfer learning to the TEM problem, we need: a pretrained neural network to use as backbone (feature extraction) and a second network to train a small, labelled TEM dataset.
- Dataset
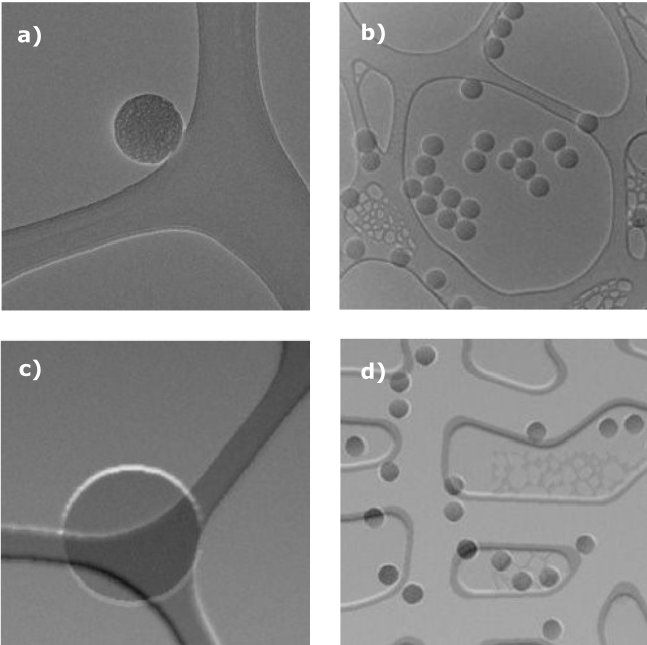
Figure 1 – Examples of (a & b) experimental data and (c & d) synthetic data.
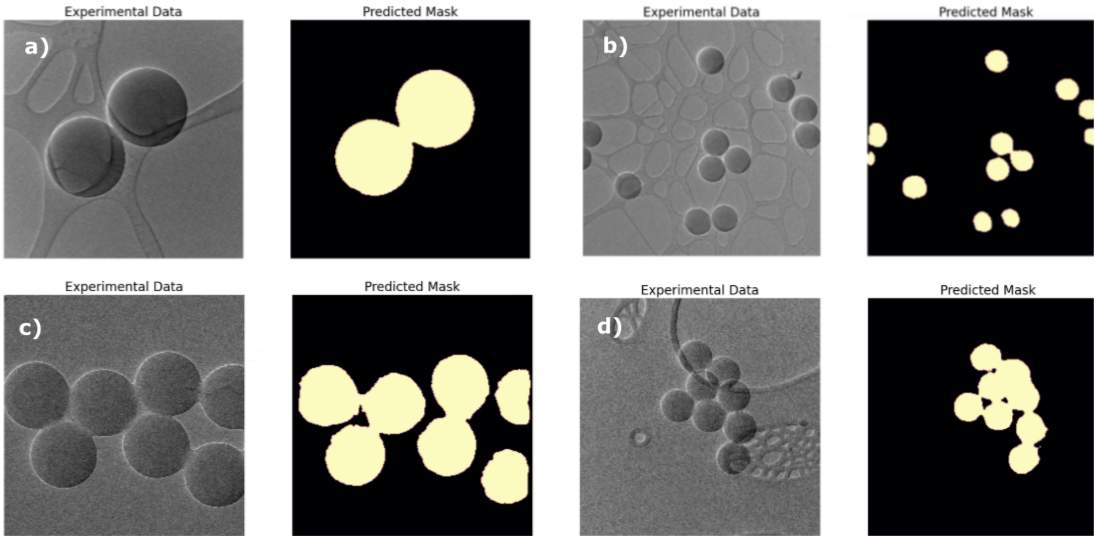
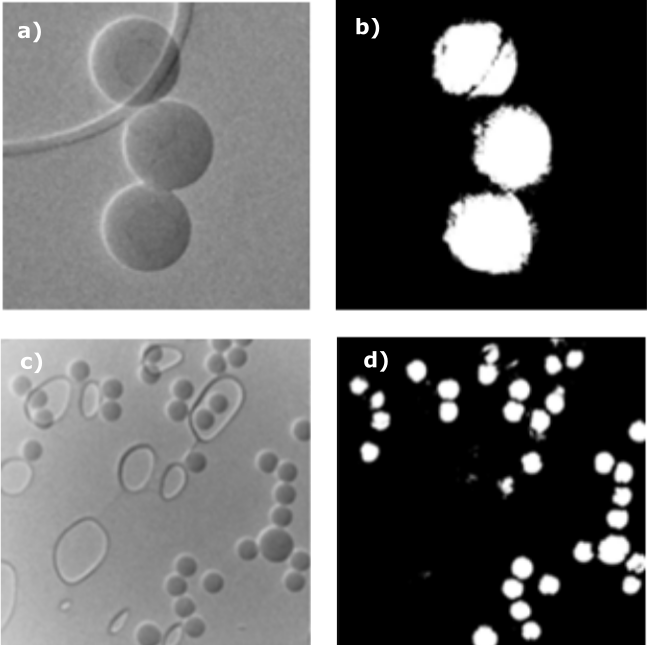
Figure 2 – Examples of the labelled data, used for training the transfer learning algorithm.
- Neural Network backbone
- Neural Network for feature extraction
Results and Discussion
We can visualise the behaviour of the algorithm in the validation set and in the sample of 11 unlabelled images which had not been used in the training were used to test the validity of the new algorithm.
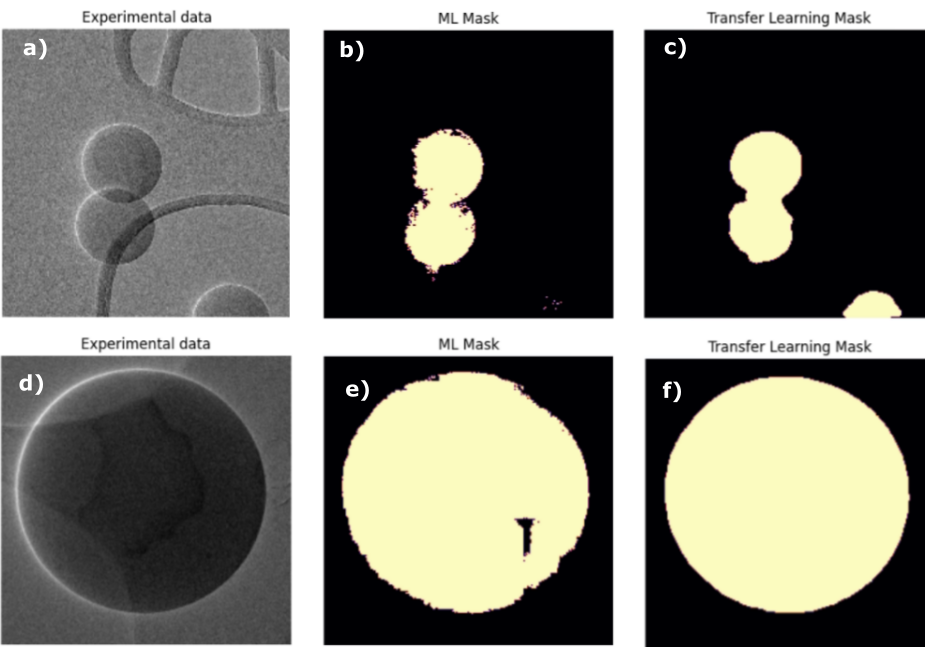
Figure 3 – Examples of (a & d) experimental data, (b & e) Machine learning labels and (c & f) transfer learning predictions in the validation set.
Figure 4 – Examples of predictions in previously unseen data (test data set).
In figure 3 we can notice that even when we have inexact masks notations, the transfer learning algorithm succeeds in segmenting the particle and its rounded shape, probably due to the weights learned with MobileNet. The network succeeds in detecting all particles in the 10 images test set, some examples are shown on figure 4.
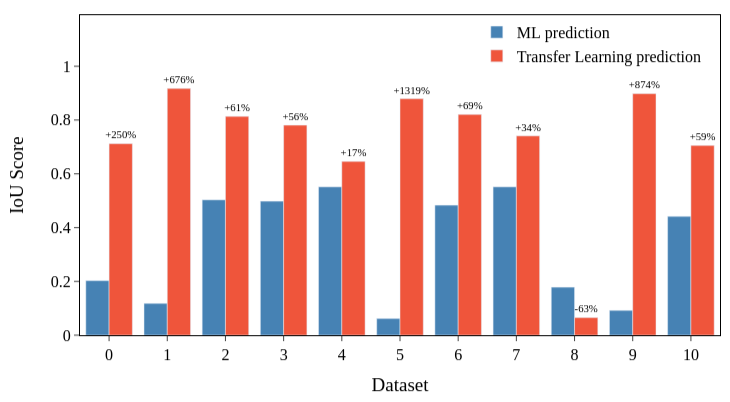
One final analysis was made using the IoU Score, which is the ratio between the true positives over the sum of true positives, false positives and false negatives and gives us a quantification of the overlap between the ground truth and the model's prediction. For the following calculations, we only consider the test set of our TEM images that were hand annotated to generate the ground truth labels. The IoU score was calculated for ground truth and machine learning prediction and for ground truth and the transfer learning model. Figure 5 show us that the proposed model correctly segments the images when ML fails.
Figure 5 - IoU Score calculated for ML and TL predictions using the ground truth test dataset.
Conclusion
We have demonstrated the use of transfer learning algorithm for image segmentation of TEM data, successfully segmenting particles from the background substrate. This method requires only a small dataset of images for training which is often the bottleneck from TEM applications in conjunction with machine learning. Further developments of the algorithm are ongoing especially the refinement of labelling used for training, via manual annotations, and building an auxiliary algorithm to identify the particles physical dimensions and morphologies automatically.
Acknowledgements
This abstract has emanated from research conducted with the financial support of Science Foundation Ireland under grant no. 18/CRT/6049.
- References
[1] Lambda X-ray Detectors, https://x-spectrum.de/products/lambda/
[2] Mrinal R. Bachute, Javed M. Subhedar, Autonomous Driving Architectures: Insights of Machine Learning and Deep Learning Algorithms, Machine Learning with Applications, Volume 6, 2021, 100164, ISSN 2666-8270, https://doi.org/10.1016/j.mlwa.2021.100164.
[3] J. Deng, W. Dong, R. Socher, L. -J. Li, Kai Li and Li Fei-Fei, "ImageNet: A large-scale hierarchical image database," 2009 IEEE Conference on Computer Vision and Pattern Recognition, Miami, FL, USA, 2009, pp. 248-255, doi: 10.1109/CVPR.2009.5206848.
[4] Image Classification on ImageNet, https://paperswithcode.com/sota/image-classification-on-imagenet.
[5] Padmavathi Kora, Chui Ping Ooi, Oliver Faust, U. Raghavendra, Anjan Gudigar, Wai Yee Chan, K. Meenakshi, K. Swaraja, Pawel Plawiak, U. Rajendra Acharya, Transfer learning techniques for medical image analysis: A review, Biocybernetics and Biomedical Engineering, Volume 42, Issue 1, 2022, Pages 79-107, ISSN 0208-5216, https://doi.org/10.1016/j.bbe.2021.11.004.
[6] Howard, Andrew G., et al. "Mobilenets: Efficient convolutional neural networks for mobile vision applications." arXiv preprint arXiv:1704.04861 (2017).
[7] Ronneberger, Olaf, Philipp Fischer, and Thomas Brox. "U-net: Convolutional networks for biomedical image segmentation." Medical Image Computing and Computer-Assisted Intervention–MICCAI 2015: 18th International Conference, Munich, Germany, October 5-9, 2015, Proceedings, Part III 18. Springer International Publishing, 2015.
[8] Du, Getao, et al. "Medical image segmentation based on u-net: A review." Journal of Imaging Science and Technology (2020).