Tomographic Reconstruction of a Twisted Pharmaceutical Cocrystal using Scanning Electron Diffraction
- Abstract number
- 36
- Presentation Form
- Contributed Talk
- DOI
- 10.22443/rms.mmc2023.36
- Corresponding Email
- [email protected]
- Session
- EMAG - Bio, Cryo & Low-dose EM Imaging
- Authors
- Joonatan Laulainen (1), Duncan Johnstone (1), Paul Midgley (1)
- Affiliations
-
1. University of Cambridge
- Keywords
Scanning Electron Diffraction (SED), 4D-STEM, Tomography, Organic, TEM, crystallography
- Abstract text
A novel twisting polymorph of a paracetamol-theophylline cocrystal was identified using transmission electron microscopy. The crystal’s orientation changed rapidly, but by using scanning electron diffraction, we were able to identify individual patterns, simulate their relative orientation, and reconstruct a 3D reciprocal space unit cell. This unit cell is distinct from the reported crystal structure and showcases nanoscale twisting.
Pharmaceutical cocrystals are a potential solution to poor solubility that plagues many pharmaceutical compounds and prevents their access to market [1]. Aqueous solubility, in particular, remains an all-too-frequent hurdle for new pharmaceutical compounds, with as many as 90% of new chemical entities suffering from poor aqueous solubility [2]. The ability to design specific cocrystals to enable nanoscale solubility, hydration properties, stability. However, such cocrystals are often difficult to characterise, due to complex phase polymorphism that is dependent on composition and processing conditions. 3D-electron diffraction (3D-ED) has been used to probe the crystal structure of such nanoscale volumes, but significant questions remain about the microstructure of pharmaceutical cocrystals.
Here, we use scanning electron diffraction (SED) [3], a 4D-STEM technique with a low (< 1 mrad) convergence angle to probe the nanostructure of a paracetamol-theophylline cocrystal. The cocrystal was prepared by ball-milling a 50%:50% mixture of off-the-shelf pure paracetamol and pure theophylline, to mimic the mechanochemical transformations that can occur in drug processing in the pharmaceutical industry. SED data was acquired on a JEOL ARM300F STEM operated at 200 kV with a beam diameter of ca. 5 nm and a convergence angle of ca. 0.6 mrad. The current was ca. 1.5 pA, and the data was acquired on a Merlin Medipix3 with a frame time of 1 ms, leading to an electron fluence of ca. 5 e-/Å2.
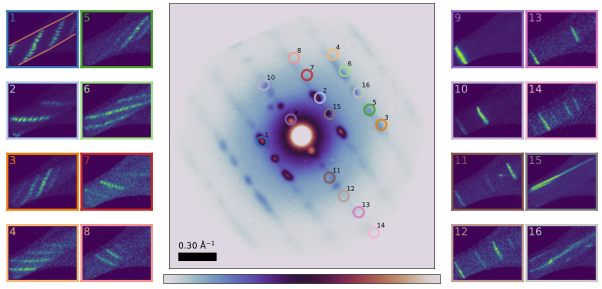
The SED data of the cocrystal showcased a previously unknown twisting polymorph in addition to the reported cocrystal structure (KIGLUI). The summed diffraction pattern showed fibre-like characteristics (Figure 1 centre), such that 3D-ED could not be used to characterise its unit cell. The twisting was identified through virtual dark field (VDF) imaging, shown in Figure 1. Twisting is characterised by multiple rotations about different axes, seen notably for the equatorial reflections (Fig 1 insets 9-14) and the perpendicular meridian reflections (insets 15-16), which showcase perpendicular VDF contrast. No such twisting was observed in similarly processed pure paracetamol or pure theophylline. A twist rate for the crystal was identified using two independent methods, by analysing the real-space displacement of the intensities of +g and -g reflections, and by analysing the relative positions of the equatorial reflections. The rotation rate was found to be similar for both methods (0.049° nm-1 and 0.058° nm-1). This rotation rate was then used to simulate a twisted object, enabling classification of the relative orientation of each observed diffraction, without knowledge of the crystallographic axes.
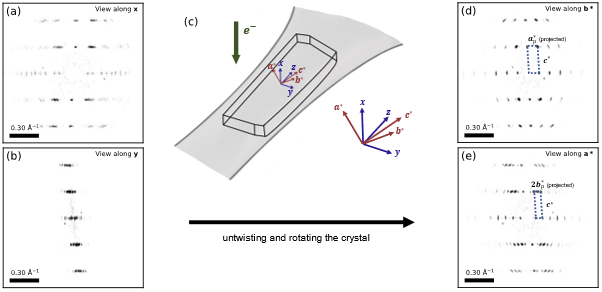
As a result, it was possible to untwist the crystal, by placing these oriented diffraction patterns in 3D reciprocal space at the appropriate relative orientation. The resulting 3D reciprocal lattice is shown in Figure 2. Diffraction spots from across the specimen lined up in a consistent lattice, and each spot in this 3D reciprocal lattice was found to correspond to exactly one band of VDF intensity. In addition, several systematic absences were identified. The 0k0 reflections (where k is odd) and 00l reflections (where l is odd) are missing. This, with the measured unit cell angles implies the twisted crystal has a monoclinic unit cell (unique c-axis chosen due to the twist orientation) with a=9.2 Å, b=26.7 Å, c=3.72 Å and a monoclinic angle γ of 98°, and the P21/b (14) space group. This unit cell was confirmed by other twisting crystals in the same data set.
In conclusion, a novel twisting polymorph for the paracetamol-theophylline cocrystal system was identified. By using SED, we were able to isolate diffraction patterns at different spatial locations across the sample, simulate their orientation, and reassemble the 3D reciprocal unit cell. This enabled us to solve the unit cell of the polymorph without prior knowledge of its structure or orientation relative to the electron beam [4].
Figure 1. Summed diffraction pattern of the particle (centre). The rod-like particle shape is highlighted in red lines in inset 1. Insets 1-16: VDF contrast of different reflection overlaid on the central pattern. Multiple diffracted spots are overlaid in some of the reflections, each resulting in its own line (such as in inset 4).
Figure 2. (a-b) Projections of the 3D reciprocal space along the beam path and perpendicular to it. (c) Schematic of the approximate crystal orientation and difference in the axes for (a-b) and (d-e). (d-e) Untwisted and rotated 3D-reciprocal space, projected along the reciprocal crystal axes b* and a*.
- References
[1] N. Qiao, M. Li, W. Schlindwein, N. Malek, A. Davies, and G. Trappitt, “Pharmaceutical cocrystals: an overview,” Int. J. Pharm., vol. 419, no. 1–2, pp. 1–11, 2011.
[2] S. Kalepu and V. Nekkanti, “Insoluble drug delivery strategies: review of recent advances and business prospects,” Acta Pharm. Sin. B, vol. 5, no. 5, pp. 442–453, 2015.
[3] C. Ophus. "Four-dimensional scanning transmission electron microscopy (4D-STEM): From scanning nanodiffraction to ptychography and beyond." Microscopy and Microanalysis 25.3 (2019): 563-582.
[4] The authors acknowledge funding from the EPSRC through doctoral training award EP/R513180/1. We thank Diamond Light Source Ltd. for access and support in the use of tmhe electron Physical Science Imaging Centre (EM16983, EM17991, EM18488, EM20527). PA.M. thanks the EPSRC for financial support under grant EP/R008779/1 and EP/V007785/1, as well as the EC H2020 grant ESTEEM3 (agreement ID 823717).