Atomic-Scale Characterisation of Structured Water Molecules at the Interface of Lipid Membranes
- Abstract number
- 70
- Presentation Form
- Poster
- Corresponding Email
- [email protected]
- Session
- Poster Session Two
- Authors
- Dr. Simone Benaglia (1, 2), Harriet Read (1, 2), Dr. Laura Fumagalli (1, 2)
- Affiliations
-
1. The University of Manchester
2. National Graphene Institute
- Keywords
3D Atomic Force MIcroscopy, Solid-Liquid Interface, lipid biophysics
- Abstract text
Water molecules are decisive in the structure and function of biomolecules and their assembly.1 Hydration is essential in biomembranes and its variation is known to regulate their intermolecular and intermembrane interactions whilst also contributing to the adaptation of their characteristics to new functional requirements. For instance, it is known that different levels of hydration change the thermodynamics of lipid bilayers, e.g. their transition temperature, enthalpy, and entropy. These properties in turn regulate important physical properties of the membranes, such as their permeability, signal propagation through the membrane, and phenomena of endocytosis and exocytosis.2 Although, there have been many scientific studies on biomembrane hydration, little is known about the local nanoscale properties of their solid-liquid interface.
To address this issue, we used three-dimensional Atomic Force Microscopy (3D AFM) to map the organisation of water molecules at the interface with phosphatidylcholine lipid membranes. 3D AFM allows probing the solid-liquid interfacial region with atomic scale resolution. The AFM tip records a change in force due to the density variation of the liquid molecules under the tip apex. 3D AFM has been mostly applied to flat, stiff, and crystalline surfaces so far.3 This is because the solid surface greatly facilitates probing the interfacial water structure. Its application on biological materials is much more challenging due to the soft nature of the surface, although recent studies have already shown great potential in mapping soft matter-water interfaces, such as those of proteins and lipids.4,5 Building on these studies, here we experimentally investigated the structural arrangement of water molecules at lipids membranes. First, we show that by carefully tuning the AFM parameters, we could probe the water-lipid interface without perturbing it. We then carried out experiments under controlled temperature above and below the lipid phase transition. We mapped the water arrangement over the lipids in their liquid and gel phase, respectively, and compared their hydration structures with 3D atomic-scale resolution. We show the interface organises in multiple hydration layers near the soft hydrophilic lipid heads and highlight the differences between the two phases.
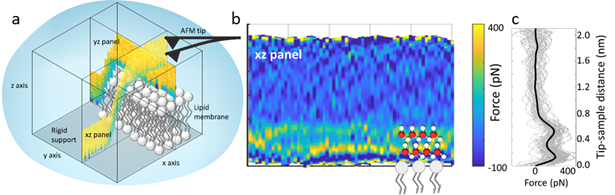
Figure: (a) Schematic of 3D-AFM on a lipid membrane. A 3D data cube is obtained (xz and yz panel are shown). (b) Example of xz force panel extracted from the 3D data cube, showing two hydration layers on a DMPC lipid membrane. (c) Force-distance curve extracted from (b). The two peaks in the force plot correspond to the positions of the two hydration layers. The black curve is the average curve.
- References
1. M. C. Bellissent-Funel, A. Hassanali, M. Havenith, R. Henchman, P. Pohl, F. Sterpone, D. Van Der Spoel, Y. Xu and A. E. Garcia, Chem. Rev., 2016, 116, 7673–7697.
2.T. Heimburg, Thermal Biophysics of Membranes, Wiley-VCH Verlag GmbH & Co. KGaA, 2007.
3. T. Fukuma and R. Garcia, ACS Nano, 2018, 12, 11785–11797.
4. E. T. Herruzo, H. Asakawa, T. Fukuma and R. Garcia, Nanoscale, 2013, 5, 2678–2685.
5. H. Asakawa, S. Yoshioka, K. I. Nishimura and T. Fukuma, ACS Nano, 2012, 6, 9013–9020.