Multispectral live cell imaging with uncompromised temporal resolution
- Abstract number
- 61
- Presentation Form
- Contributed Talk
- DOI
- 10.22443/rms.mmc2023.61
- Corresponding Email
- [email protected]
- Session
- Optical Imaging of Fast Dynamic Processes
- Authors
- Akaash Kumar (1, 2), Emmanuel Derivery (1), James Manton (1)
- Affiliations
-
1. MRC Laboratory of Molecular Biology
2. University of Cambridge
- Keywords
Live Cell Imaging
Light Sheet Microscopy
Multispectral Imaging
Cellular Dynamics
- Abstract text
The hallmark of life is that it is animate. Advances in live cell fluorescence microscopy have enabled scientists to shed light on the spatiotemporal nature of biological processes, but conventional methods are temporally limited when imaging more than one fluorescent label of interest. Due to the broad emission spectra of typical fluorescent labels, it is often not possible to observe more than three or four fluorescent labels in the same sample. Spectral unmixing provides a way around this issue, by reassigning spectrally overlapping signals, but typically requires slow point-scanning acquisition and high signal levels, incompatible with live-cell imaging.
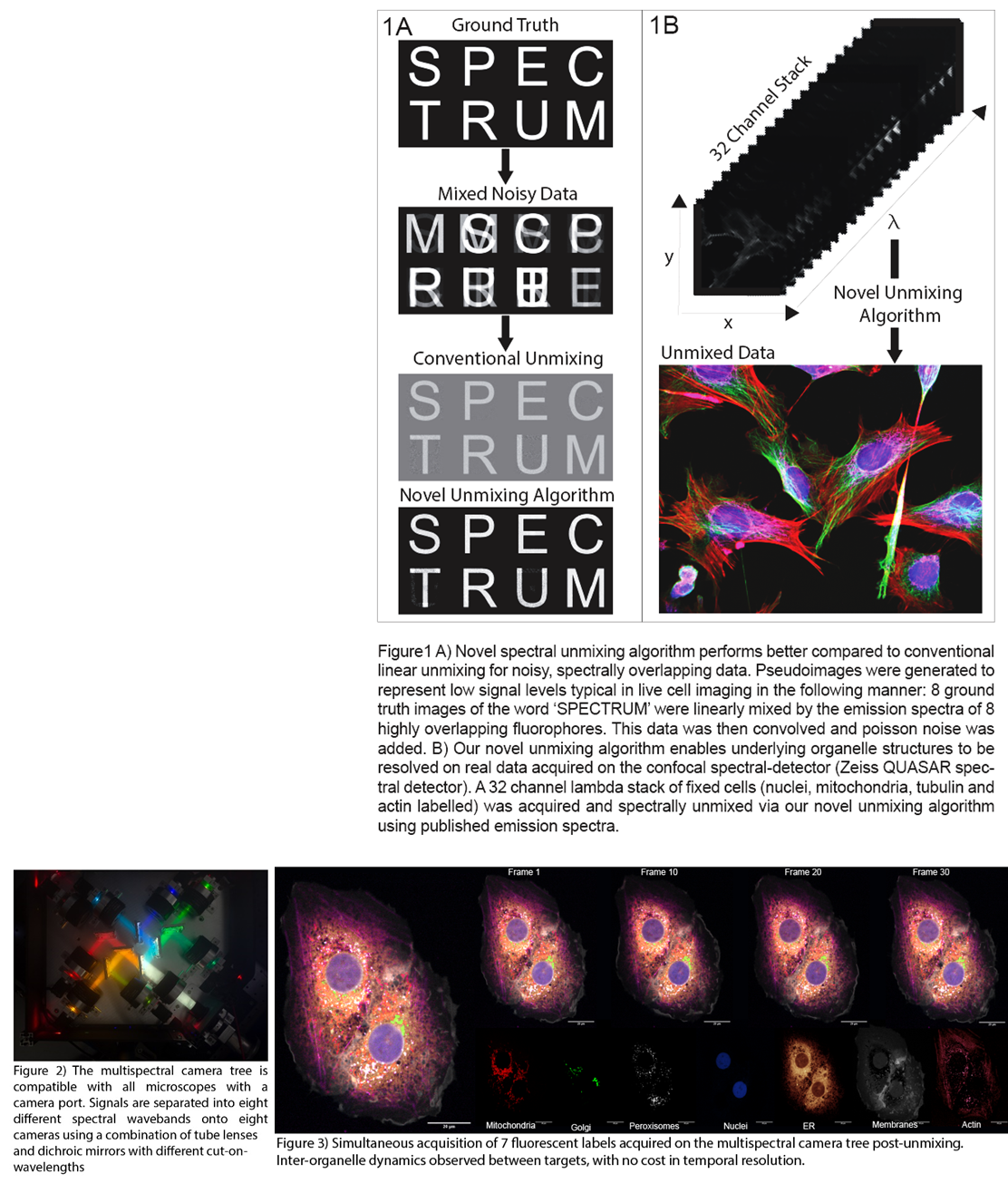
Here, we first introduce our newly developed iterative spectral unmixing algorithm, specifically designed to perform well with the low signal levels typically found in live cell imaging. We demonstrate the superiority of our algorithm for unmixing noisy data compared to conventional linear unmixing (Fig1A) and show its application to spectral-detector confocal imaging data (e.g. as produced by Zeiss QUASAR) (Fig1B). We then describe the design of our novel multispectral imaging system (Fig2), which is capable of acquiring spectrally-resolved, video-rate data across the entire field of view simultaneously, and show how it can be combined with any camera-based microscope. We use our camera system and unmixing software with spinning disk confocal and light sheet fluorescence microscopy to demonstrate the rapid live-cell imaging of >6 fluorescent labels with high photon efficiency and minimal sample perturbation (Fig3). We conclude with an outlook on expanding the number of fluorescent labels further still, exploiting NIR fluorophores, and apply our methods to questions in endosomal sorting and membrane remodelling.