Cryogenic Electron Ptychographic Single Particle Analysis (Cryo-EPty SPA) with Tunable Information Transfer
- Abstract number
- 58
- Presentation Form
- Contributed Talk
- DOI
- 10.22443/rms.mmc2023.58
- Corresponding Email
- [email protected]
- Session
- EMAG - Bio, Cryo & Low-dose EM Imaging
- Authors
- Mr Xudong Pei (1), Dr Liqi Zhou (1), Dr Chen Huang (6), Dr Mark Boyce (4), Dr Judy S. Kim (6, 2), Dr Emanuela Liberti (6), Dr Takeo Sasaki (5), Prof Peter D. Nellist (2), Prof Peijun Zhang (4, 7), Prof David I. Stuart (4), Prof Angus I. Kirkland (6, 2), Dr Peng Wang (3)
- Affiliations
-
1. College of Engineering and Applied Sciences, Nanjing University
2. Department of Materials, University of Oxford
3. Department of Physics, University of Warwick
4. Division of Structural Biology, Welcome Trust Centre for Human Genetics, University of Oxford
5. JEOL Ltd., 3-1-2 Musashino, Akishima
6. The Rosalind Franklin Institute, Harwell Science and Innovation Campus
7. Electron Bio-Imaging Centre, Diamond Light Source, Harwell Science and Innovation Campus
- Keywords
Cryo-EM, Ptychography, SPA, Phase Imaging
- Abstract text
In this talk, we will propose a new 3D SPA technique based upon cryogenic ptychography (cryo-EPty SPA) and experimentally demonstrate that this technique can restore 3D information of a sample across different bandwidths of spatial frequencies by adjusting convergence semi-angle (CSA) of the electron probe.
Rapid advances in the field of cryo-electron microscopy (cryo-EM) have made it possible to visualize three-dimensional structures of a wide range of biological macromolecules at near-atomic resolution in a close-to-native, frozen-hydrated state [1]. However, the high electron beam sensitivity of vitrified biological samples [2] results in issues such as low signal-to-noise ratios (SNR) [3] and low contrast [4,5] in conventional cryo-EM images. To overcome these problems, cryo-EM single particle analysis (SPA) usually records micrographs at high defocus to improve information transfer at low spatial frequencies. However, the high-defocus condition rapidly reverses high-frequency information in the phase contrast transfer function (PCTF) [6], which causes difficulties in accurate identification, classification and alignment of particles to a common reference.
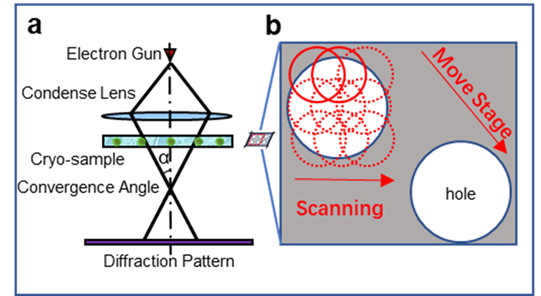
An alternative strategy is based on scanning ptychographic diffractive imaging using phase information recovered by electron ptychography [7] as shown in Fig. 1a, which has been recently demonstrated under low-dose conditions for biological samples embedded in vitreous ice [8]. Electron ptychography has generated considerable interest given its potential for super-resolution imaging [9,10], high-contrast light-element detection [11], low-dose imaging [12] and three-dimensional imaging [13,14]. Moreover, as ptychography utilizes the full diffraction pattern, it is dose-efficient particularly when data is recorded using direct electron detectors which record data with high signal-to-noise at low electron dose. This approach has recently been demonstrated for micrometer-wide phase reconstruction of an unstained virus-infected cell at a dose of 27 e/Å2 [8].
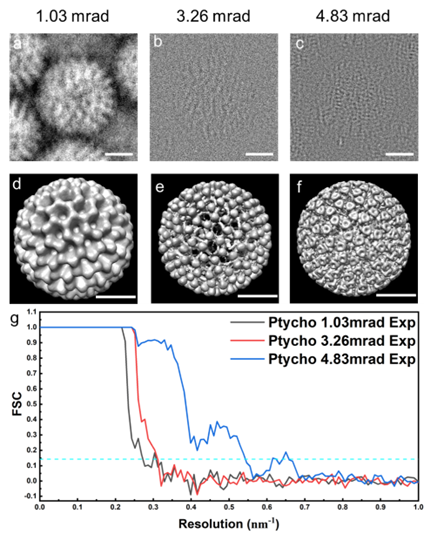
Experimental cryo-EPty SPA datasets (Fig. 1 b) were acquired in a scanning diffraction configuration, in which a defocused probe is scanned over a cryo-sample. Using the ePIE algorithm [8], the ptychographic phase of rotavirus double-layered particles (DLPs) with 76.5 nm in diameter were reconstructed at doses of 22.6, 24.1 and 24.6 e/A2 corresponding to CSA of 1.03, 3.26, and 4.83 mrad as shown in Fig. 2a-c. We subsequently use a standard SPA pipeline to reconstruct 3D density maps (Fig. 2d-f) of rotavirus DLPs with around 300 particles from the stack of particle phases [7]. The resolution of the 3D maps is estimated as 3.72 nm, 3.29 nm, and 1.86 nm, respectively using a Fourier Shell Correlation (FSC=0.143, gold-standard) as shown in Fig. 2g. We expect that cryo-EPty combined with SPA has great potential to yield high-resolution 3D reconstructions of biological samples [16].
Figure 1 | Schematic diagrams of the optical configuration (a) and data acquisition (b) for ptychography
Figure 2 | 3D Rotavirus DLPs reconstructions using Cryo EPty-SPA for various CSAs. (a to c) Experimental ptychographic phase of a virus particle with CSAs, a = 1.03 mrad, 3.26 mrad and 4.83 mrad, respectively. (d to e) 3D maps corresponding to the particle instances (a to e). (g) Fourier shell correlation curves of the 3D maps in (d to f) Scale bars: 25 nm.
- References
References:
[1] R Henderson et al., Journal of Molecular Biology 213 (1990), p. 899.
[2] KA Taylor et al., Science 186 (1974), 1036.
[3] R Henderson, Quarterly Reviews of Biophysics 28 (2009), p. 171.
[4] R Danev et al., Ultramicroscopy 88 (2001), p. 243.
[5] R Danev et al., Proceedings of the National Academy of Sciences 111 (2014), p. 15635.
[6] K H. Downing et al., Ultramicroscopy 108 (2008), p. 921.
[7] JM Rodenburg, Advances in Imaging and Electron Physics 150 (2008), p. 87.
[8] L Zhou, et al., Nature Communications 11 (2020), p. 2773.
[9] PD Nellist et al., Nature 374 (1995), p. 630.
[10] Y Jiang et al., Nature 559 (2018), p. 343.
[11] P Wang et al., Scientific Reports 7 (2017), p. 2857.
[12] J Song et al., Scientific Reports 9 (2019), p. 3919.
[13] AM Maiden et al., J. Opt. Soc. Am. A 29 (2012), p. 1606.
[14] S Gao et al., Nature Communications 8 (2017), p. 163.
[15] AM Maiden et al., Ultramicroscopy 109 (2009), p. 1256.
[16] The authors acknowledge funding from University of Warwick Research Development Fund (RDF) 2021-22 Science Development Award