Counting the cold: do flowering plants encode winter memory with large nuclear assemblies?
- Abstract number
- 107
- Presentation Form
- Contributed Talk
- DOI
- 10.22443/rms.mmc2023.107
- Corresponding Email
- [email protected]
- Session
- Optical Imaging of Fast Dynamic Processes
- Authors
- Dr Alex Payne-Dwyer (2), Dr Govind Menon (1), Dr Geng-Jen Jang (1), Prof. Dame Caroline Dean (1), Prof Martin Howard (1), Prof. Mark Leake (2)
- Affiliations
-
1. John Innes Centre
2. University of York
- Keywords
Epigenetic memory
Protein assembly
Molecular counting
Single molecule imaging
Diffusion
Plant imaging
- Abstract text
Canonical epigenetic regulation involves addition of chemical marks, such as trimethylation, to histones. These marks encode silenced genetic states and maintain functional memory and state switching in cells. However, the small number of marks at each locus means this process alone is insufficient to retain memory over multiple cell cycles. We consider a complementary framework [1] in which large protein assemblies can provide the robust feedback needed to avoid degradation of memory, and demonstrate rapid single-molecule imaging to support this model.
Silencing of the gene FLOWERING LOCUS C (FLC) in Arabidopsis thaliana directly regulates timing of flowering after winter [2]. We adapt single-molecule microscopy [3] to the SlimVar technique for imaging live root tips. We then track the in vivo dynamics of the Polycomb-interacting protein, VRN5, and its counterpart VIN3, which is induced at low temperatures [4].
We find that these proteins form striking, dynamic assemblies of tens of molecules in the nucleus in response to prolonged cold exposure. These assemblies are sufficiently large and persistent to act as strong candidates in our model for initiating and maintaining the silenced state of the FLC gene, leading to vernal flowering. Our model indicates that the robust statistics of self-assembly could have important, general implications for epigenetic processes in eukaryotes.
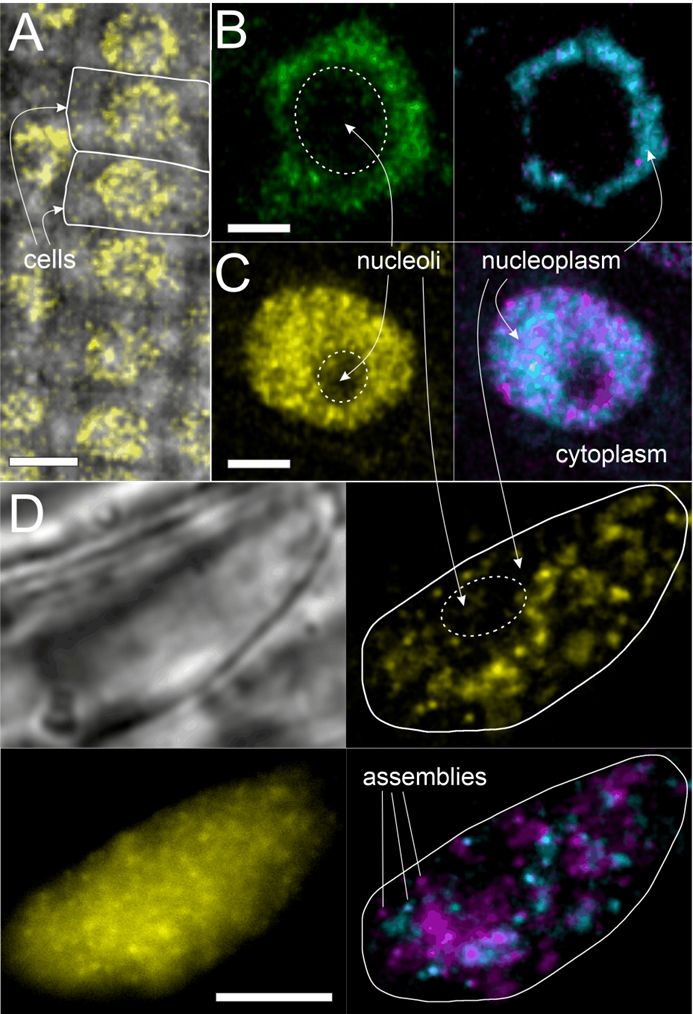
Figure. SlimVar resolves single particle dynamics of VIN3 and VRN5 in live root tips. A) Confocal image of VRN5, scale bar 10 µm. B,C) Airyscan images of VIN3 and VRN5 after cold exposure showing assemblies. Dynamics represented by median (cyan) and standard deviation (magenta) of intensity over ~100 ms exposure, indicating faster detected motion of VRN5 compared to VIN3 assemblies; scale bars 2 µm. D) SlimVar microscopy of VRN5 reveals distinct assemblies with differential mobility on the timescale of ~10 ms; scale bar 5 µm.
- References
[1] Lövkvist, C. et al. (2021) eLife 10:e66454.
[2] Costa, S. and Dean C. (2019) Biochem Soc Trans. 47 (4):1187–1196
[3] Payne-Dwyer A. et al. (2022) J R Soc Interface. 19(193):20220437
[4] Franco-Echevarría, E. et al. (2022) J Biol Chem. 298(11):102540