Automated Segmentation of Mitochondria in Virus-Infected Cells using Deep Learning Models
- Abstract number
- 227
- Presentation Form
- Poster & Flash Talk
- DOI
- 10.22443/rms.mmc2023.227
- Corresponding Email
- [email protected]
- Session
- EMAG - EM Data Processing & Analysis
- Authors
- Dr. Katherine Lau (1), Matthijs Schrage (1), Mario-Alin Rus (1), Dr. Thomas Burgoyne (3), Marre Niessen (1), Andreia Pinto (2)
- Affiliations
-
1. Delmic B.V.
2. Royal Brompton Hospital
3. University College London
- Keywords
- Abstract text
Summary
An automated mitochondrial segmentation model is being developed to facilitate the quantitative analyses of electron microscopy (EM) data of viral-infected cells. The segmentation accuracies of deep learning (DL) models MitoNet and U-Net were assessed through their Intersection over Union (IoU) score. MitoNet initially did not segment mitochondria with unhealthy morphologies well. By fine-tuning the MitoNet model, it was possible to increase the segmentation accuracy even for the diseased mitochondria. Our results demonstrate the potential of improving the pre-trained generalised models with new data to make it suitable for infectious disease research.
Introduction
Infections in cells often manifest in abnormal organelle morphologies, which can be studied by EM thanks to its high resolution. EM analysis of organelles can increase our understanding of how the infections impact the cells. Using a high speed scanning electron microscope (SEM) such as FAST-EM (Delmic B.V., Netherlands), it is furthermore possible to accelerate research. However with the large data throughput, it is necessary to reliably automate the organelle segmentation to allow for meaningful data analyses. Thus, we aim to develop a DL model to auto-segment organelles in SARS-CoV-2 infected cells. We start with mitochondria of which abnormalities such as swelling and disorganised cristae have been observed in diseased cells [1] and in SARS-CoV-2 infected cells. There is already a generalised DL model MitoNet based on DeepLabV3+ [2] available online, which has been pre-trained on mitochondria from different cell lines and is implemented on the Napari platform for users with little to no experience with DL. It was demonstrated to segment mitochondria from cell types outside its training data [4] .There is another easily accessible DL model called U-Net [3]. Attention U-Net can be trained on ZeroCostDL4Mic [5], which provides a user-friendly interface built on Google Colab for those with some programming experience but without extensive experience in DL. We compare the functionality of these tools for mitochondrial segmentation in infected cells.
Methods
Whole sections of human airway epithelial cells infected by SARS-CoV-2 Delta variant were imaged using FAST-EM at a 4 nm resolution and a 10 μs dwell time. Abnormal mitochondria in images cropped to 512x512 were annotated to train DL models and for assessing the segmentation accuracies against ground truth by calculating the IoU score, a commonly used metric for evaluating performance of deep segmentation networks [6]. We compared an Attention U-Net model (based on scripts from ZeroCostDl4Mic) trained on our annotated data, the original MitoNet and MitoNet fine-tuned with our annotated data.
Results and Discussion
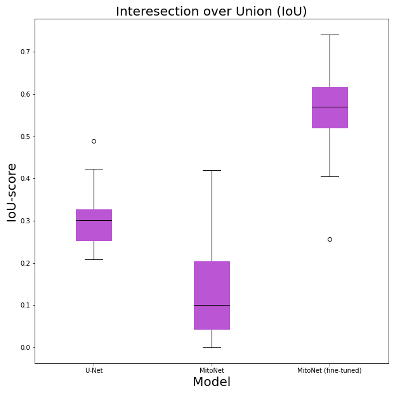
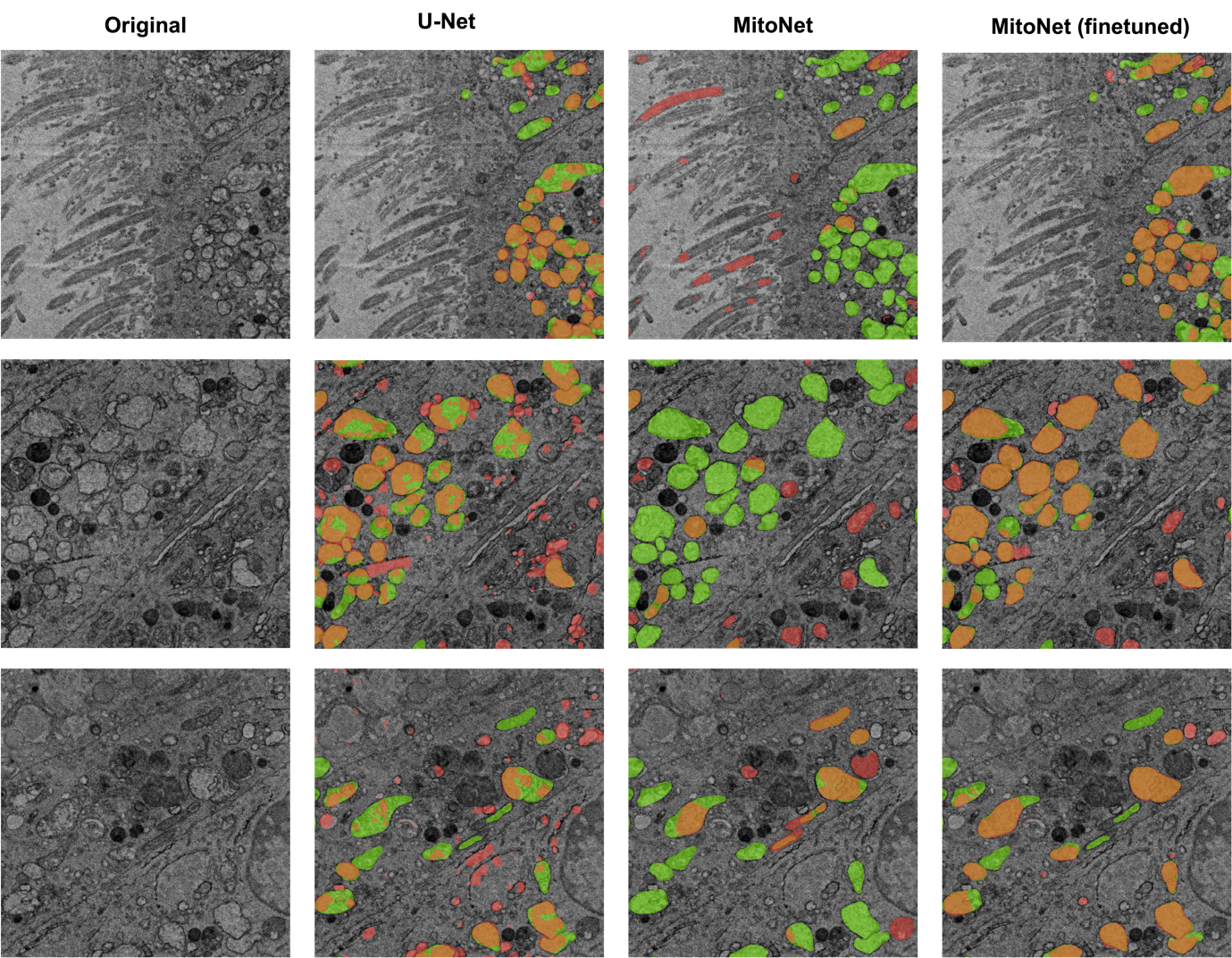
The original MitoNet and the U-Net trained on our annotated data obtained IoU scores below 0.3. The fine-tuned MitoNet model outperformed the U-Net 2-fold and the original MitoNet 5-fold (Figure 1). The original MitoNet tended to result in false negatives for diseased mitochondria (Figure 2) presumably due to MitoNet being trained on healthy mitochondria. The fine-tuned MitoNet identified almost all of the diseased mitochondria and produced fewer false positives. Additionally, MitoNet produced less fragmented segmentations compared to U-Net, possibly due to differences in architecture between DeeplabV3+ and U-net. DeeplabV3+ contains atrous convolutions to aggregate context around features.
Figure 1. IoU scores of U-Net, MitoNet, fine-tuned MitoNet on a mitochondria segmentation task in SARS-CoV-2 infected cells.
Figure 2. Left column: without annotation. Right three columns: segmentation by U-Net, MitoNet, fine-tuned MitoNet against ground truth. Orange: true positives. Green: false negatives. Red: false positives.
Although the original MitoNet did not perform well initially, it was possible to boost the performance through retraining the model with our data. On the other hand, training a mitochondrial segmentation U-Net from scratch resulted in lower IoU-scores, suggesting an advantage to work with a pre-trained generalised model. We aim to further improve its accuracy through i) increasing the training data volume and ii) 3D segmentation, as mitochondria may be more distinguishable in 3D. Developing a model that can achieve an accuracy >75% in segmenting mitochondria in SARS-CoV-2 infected cells will allow us to study the effects of different SARS-CoV-2 variants through quantitative data analysis.
Large-scale Open-source datasets and collaborative large-scale annotation in recent years have made it possible for models such as MitoNet to be integrated into the pipeline of researchers without extensive knowledge of DL. We believe generalised DL models for other organelles will greatly facilitate quantitative analysis in cellular research. Generalised organelle segmentation models require large training datasets from different cell types. High speed EM imaging technologies such as the FAST-EM can allow more training data to be acquired from different cell types within a shorter time.
Conclusion
The generalised organelle segmentation DL model MitoNet initially segmented mitochondria poorly in SARS-CoV-2 infected cells. Re-training the model with our data including diseased mitochondria greatly enhanced its performance. Further improvements to bring the segmentation accuracy to >75% will facilitate the study of mitochondria from cells infected by different SARS-CoV-2 variants. The development of generalised DL models for other organelles will accelerate the process of obtaining reliable auto-segmentation models and benefit cell biology research.
- References
- Shami, G.J., Cheng, D., Verhaegh, P. et al (2021). Three Dimensional Ultrastructure of Giant Mitochondria in Human Non-Alcoholic Fatty Liver Disease. Scientific Reports 11:3319
- Chen, L. C., Papandreou, G., Kokkinos, I. et al (2018). DeepLab: Semantic Image Segmentation with Deep Convolutional Nets, Atrous Convolution, and Fully Connected CRFs. IEEE Transactions on Pattern Analysis and Machine Intelligence, 40(4), 834-848.
- Ronneberger, O., Fischer, P., & Brox, T. (2015). U-Net: Convolutional Networks for Biomedical Image Segmentation. In International Conference on Medical image computing and computer-assisted intervention (pp. 234-241).
- Conrad, R., & Narayan, K. (2023). Instance segmentation of mitochondria in electron microscopy images with a generalist deep learning model trained on a diverse dataset. Cell Systems, 14(1), 58-71.
- von Chamier, L., Laine, R. F., Jukkala, J., Spahn, C., Krentzel, D., Nehme, E., ... & Henriques, R. (2021). Democratising deep learning for microscopy with ZeroCostDL4Mic. Nature communications, 12(1), 2276.
- He, K., Gkioxari, G., Dollár, P., & Girshick, R. (2017). Mask r-cnn. In Proceedings of the IEEE international conference on computer vision (pp. 2961-2969).