Tiling-based Multislice for Transmission Electron Microscopy
- Abstract number
- 338
- Presentation Form
- Poster & Flash Talk
- DOI
- 10.22443/rms.mmc2023.338
- Corresponding Email
- [email protected]
- Session
- EMAG - EM Data Processing & Analysis
- Authors
- Mr Stephen Mullins (2, 3), Dr Andrew Stewart (1)
- Affiliations
-
1. Department of Chemistry, University College London
2. Department of Physics, University of Limerick
3. SFI Centre for Research Training in Foundations of Data Science, University of Limerick
- Keywords
simulation, multislice
- Abstract text
Multislice is an algorithm which simulates dynamical electron scattering as an electron beam passes through a sample model to simulate an image, and is calculated by splitting the model into sufficiently thin slices to ensure kinematic electron scattering within each layer, finally producing an exit wave as the propagation exits the bottom of the specimen [1]. There are various multislice algorithms available for Transmission Electron Microscopy (TEM) image simulations, including abTEM [2], MULTEM [3], and Dr Probe [4], that are used to verify models for interpreting experimental information. However, there are frequent and similar limitations amongst these multislice programs, in particular, a narrow field of view due to the limited memory capacity in standard computers to calculate larger images in a reasonable time period. This limited field of view can affect the ability to clarify and validate experimental images with simulated images; for example, dense amorphous specimens or considerably large non-crystalline materials can be particularly challenging.
We propose to increase the simulation field of view and size of the simulated images by dividing the model into a series of smaller regions, referred to as tiles, and perform multislice on each tile of the tiles separately, generating individual images or exit waves to be merged together, with each tile being small enough to fit into the available memory and be effectively calculated via the standard multislice methodologies [5].
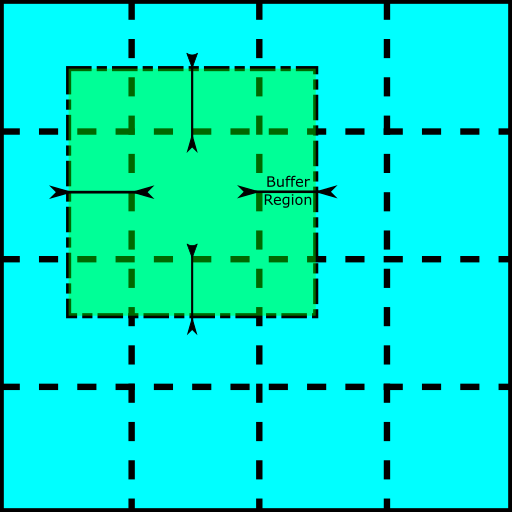
In the method presented here, each atomic layer in the model is separated into a series of square tiles, padded with a buffer region to ensure consistency and continuity between the tiles and reduce the edge effects. Next, an electron wave is generated for each tile series, and the Fast Fourier Transform (FFT) convolution is applied in parallel for each electron wave during the multislice, which is repeated until the output exit waves are determined for each tile. Finally, the buffer regions of the output exit waves are stripped away, and the remaining wave segments are assembled into a single wave function.
Here we present advances in the parallelised tiling-based multislice algorithm, which can feasibly simulate a large field of view images to help validate and interpret TEM images and exit wave functions.
Figure 1: The fragmenting of an atomic potential slice into a set of 4x4 tiles, with each tile granted a buffer region overlapping adjacent tiles.
- References
[1] Kirkland, J. (2020). Advanced Computing in Electron Microscopy (2nd edition.). Springer Publishing.
[2] Madsen, Jacob & Susi, Toma. “abTEM: ab Initio Transmission Electron Microscopy Image Simulation”. Microscopy and Microanalysis. 26. (2020).
[3] I. Lobato, D. Van Dyck, “MULTEM: A new multislice program to perform accurate and fast electron diffraction and imaging simulations using Graphics Processing Units with CUDA”, Ultramicroscopy, 156, (2015)
[4] J. Barthel, Dr. Probe: A software for high-resolution STEM image simulation, Ultramicroscopy, 193, (2018).
[5] S. Ali, M. Du, M. Adams, B. Smith, and C. Jacobsen, "Comparison of distributed memory algorithms for X-ray wave propagation in inhomogeneous media”, Opt. Express, 28, (2020).